1. Project Background
In modern bovine genetic improvement systems, in vitro fertilization (IVF) combined with embryonic genetic screening has become a core technological approach for enhancing herd genetic merit and accelerating the dissemination of superior germplasm. Conventional embryonic genetic testing methods predominantly rely on invasive biopsy procedures, which are not only technically demanding and costly, but may also compromise embryonic integrity and developmental potential, thereby adversely affecting embryo transfer success rates and pregnancy outcomes.
Meanwhile, in vitro–produced embryos exhibit a relatively high incidence of chromosomal abnormalities. In the absence of effective screening strategies, such abnormalities substantially increase the risks of embryonic developmental failure and early pregnancy loss.
Against this backdrop, the breeding industry has an urgent demand for non-invasive embryonic genetic testing technologies that are highly sensitive, capable of parallel multi-parameter analysis, and suitable for large-scale application. Nanopore sequencing–based non-invasive embryo testing, leveraging its long-read capability, real-time sequencing, and highly flexible targeted analysis, provides a scalable and industrially viable solution for bovine embryo sex determination, chromosomal karyotype analysis, and the selection of favorable genetic traits
2. Project Introduction
This project focuses on bovine blastocysts generated through in vitro fertilization (IVF). Embryo-derived cell-free DNA (cfDNA) is obtained from blastocyst culture media as well as from blastocoelic fluid, and subjected to whole-genome amplification followed by nanopore sequencing–based analysis to enable multidimensional genetic evaluation of embryos.
Nanopore sequencing generates ultra-long continuous reads that can span repetitive sequences, complex genomic regions, and large-scale copy number variation (CNV) segments. This intrinsic advantage confers superior performance in the detection of chromosomal aneuploidy and the identification of structural variants—such as large deletions, duplications, and translocations—in bovine embryos, compared with short-read next-generation sequencing platforms and microarray-based technologies. Consequently, this approach is particularly well suited for the screening of potentially lethal or developmentally restrictive chromosomal abnormalities at the embryonic stage, thereby providing a robust genetic basis for embryo selection in assisted reproductive and breeding programs.
3. Project Process




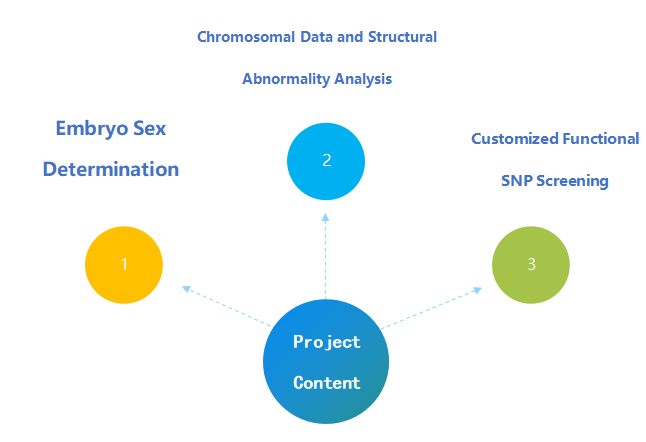
4. Project Content
5. Applicable Testing Scenarios
Small-Scale Livestock Farmers and Breeding Operations
In Vitro Fertilization (IVF) and Embryo Culture Laboratories
National or Regional Cattle Breeding Enterprises
Agricultural Research Institutes and University Research Groups
6. Project Advantage
1. Non-invasive testing significantly improves embryo utilization efficiency: By obtaining genetic information from blastocoelic fluid and cell-free DNA (cfDNA) in embryo culture media, this approach avoids the interference of conventional embryo biopsy with embryonic structure and developmental potential. Compared with invasive testing methods, it effectively reduces the risk of embryo damage and increases implantation and pregnancy success rates after blastocyst transfer, thereby fundamentally enhancing the practical utilization of IVF embryos and overall breeding efficiency.
2. Simultaneous sex determination, karyotype analysis, and multi-trait screening in a single assay: A single integrated testing workflow enables embryo sex determination, chromosomal karyotype analysis, and targeted screening of economically important SNP traits, overcoming the limitations of conventional approaches that require multiple workflows for individual indicators. This multi-trait, joint analytical strategy allows for comprehensive genetic quality assessment at the embryonic stage, thereby reducing uncertainty in subsequent breeding and husbandry practices.
3. Significant reduction in the overall cost per embryo: Non-invasive sampling minimizes embryo loss and increases the proportion of transferable embryos within a single IVF cycle, thereby improving overall resource utilization. In parallel, the integration of multi-trait analyses into a single assay eliminates the cost accumulation associated with repeated, single-parameter tests. Together, these factors facilitate economies of scale and substantially reduce the comprehensive cost of single-embryo genetic testing.
4. Strong scalability and platformization potential: Nanopore sequencing platforms feature flexible configurations and adjustable throughput, making them well suited for demand-driven testing scenarios. Compared with large, fixed-throughput sequencing systems, they require lower upfront capital investment and offer greater flexibility in per-run operating costs, facilitating distributed deployment and scalable replication in breeding centers, IVF laboratories, and related settings. Moreover, this solution can be readily extended to other livestock species (such as dairy goats, sheep, and pigs) and to a broader spectrum of genetic traits. This inherent extensibility supports continuous product iteration and platform-based development, providing a solid foundation for sustained commercial growth.